Methods

Step 1: Sample collection, dissection and fixation
We collect and fix genital tissues from fly pupae at particular time points during development:
- We collect Drosophila melanogaster male puparia and incubate them at 25˚C prior to dissection.
- We dissect pupal terminalia in phosphate buffered saline (PBS) at 28 hours or 48 hour after puparium formation and fix them in paraformaldehyde at room temperature.
- After washing, we store our samples in ethanol at -20˚C.
Download our detailed dissection protocol.
Step 2: Probe design and synthesis
We design and synthesize antisense RNA probes that will bind messenger RNA for our target genes:
- We design 200-300 base pair probes for the largest exon present in all annotated isoforms.
- We use PCR to amplify genomic sequence with an added T7 RNA polymerase binding sequence, and we synthesize our probes using in vitro transcription.
- We purify our probes using ethanol precipitation and resuspend them in 50% formamide after analysis by Nanodrop.
Download our detailed probe protocol.

Step 3: In situ hybridization
We visualize gene expression patterns by applying probes to our samples and binding an antibody conjugated to an enzyme that catalyzes dye deposition:
- We use the InsituPro VSi from Intavis (locally nicknamed Artoo Situ) to carry out in situ hybridization, although the same experiments can be performed by hand.
- We bind digoxigenin-labeled probes to target mRNA, and recognize those probes with an antibody conjugated to alkaline phosphatase.
- When exposed to additional chemicals, alkaline phosphatase catalyses the deposition of a purple dye in regions where target mRNA is expressed.
Download our detailed protocol (robot) and (by hand).
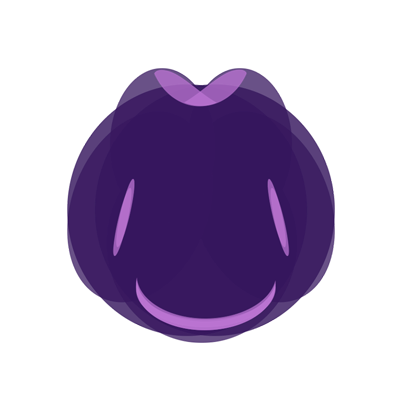

Step 4: Mounting and imaging
We image our samples using light microscopy:
- We mount samples on microscope slides coated with ploy-L-lysine to help orient them correctly.
- We image samples at 20X magnification on a Leica DM 2000 with a Leica DFC540 C camera.
- Each image was Z stack-compiled with the Leica Application Suite to allow for visualization of expression patterns at different depths in the sample.